Save As: DNA 🧬 Part 2
The Part Where Things Get Very Very Real

Welcome to part 2 of Save As: DNA. In part 1, EXO stopped off at two pivotal points in the twentieth century that put the possibilities of saving digital information in genetic material on the map. But we’re post-potential now. DNA data storage is extremely here.
In part 2, EXO goes deep into the IRL applications of DNA data storage. Here’s how it’s going to go:
Breakthroughs - A trip through recent groundbreaking achievements in DNA data storage.
DIY DNA Data Storage - A step-by-step guide to storing digital data in DNA.
Industry Forefront - An interview with world-leading synthetic DNA data storage company Twice Bioscience.
Let’s go.

Breakthroughs
While meaningful DNA data storage research has been ongoing for 40 years, major strides forward in the space really picked up in 2011. It’s difficult to identify what kicked this experimentation into high gear. The drastic decrease in decoding errors when reading data out of DNA seems to have played role. Bolstered by real promise, private and academic research groups jumped into the fray. Here are some serious wt actual f moments over the last decade:
August, 2012 - The Wyss Institute for Biologically Inspired Engineering at Harvard University teams up with George Church to encode an HTML draft of a 53,400-word book, eleven JPEG images and one JavaScript program into DNA. The team printed 70 billion copies — roughly three times the combined print run of the top 100 books of all time.
February, 2015 - Swiss researchers prove that data can live in DNA virtually rent free for 2,000 years if kept at a temperature of 9.4 degrees celsius. At -18 degrees celsius, data can be stored in DNA for 2 million years, with almost no maintenance.
December, 2016 - Dina Zielinski and Yaniv Erlich use a technique called DNA Fountain to encode a data ’tarball’ consisting of a complete graphical operating system and a $50 Amazon gift card into a DNA oligo pool (a collection of gene sequences). Using a Python script on a standard laptop, the team decodes the data with 100% accuracy in 9 minutes. The data density comes in at 215 Petabytes per gram.
June, 2019 - DNA storage platform Catalog crams the entirety of Wikipedia’s English-language version - 16 gigabytes - into a single laboratory vial of DNA.

December, 2019 - A Swiss and Israeli team encode the 3D design instructions for a bunny figurine into DNA silicate beads. The beads are then mixed into a precursor material for 3D printing. The team reads the DNA back and use it to 3D print a bunny that contains 370 million copies of its own DNA - just like a biological system. The same team also encodes a 1.4 megabyte Youtube video into DNA, mixes it into plexiglass and fabricates it into reading glasses. The spectacles containing the video look exactly like identical glasses that don’t contain DNA, suggesting potential DNA data storage information-hiding applications.
July, 2020 - Chinese scientists at Tianjin University store and retrieve 445 kilobytes of digital data from living E. coli bacteria cells. As the DNA molecules replicate so does the data, with near-perfect fidelity. The ability to extract data from an expanding pool of DNA signals high data-stability over longer periods of time, on the cheap.
August, 2020 - Twist Bioscience works with researchers at ETH Zurich to encode the 750-gigabyte first episode of the Netflix series ‘Biohackers’ into DNA nucleotides.
February, 2021 - DNA is successfully extracted from the tooth of a steppe mammoth which was been buried in Siberian permafrost for 1.2 million years, proving that DNA can preserve data for over a million years.
DIY DNA Data Storage (aka how tho)
The cutting-edge nature of DNA data storage that makes it so exciting also serves as a barrier to widespread understanding. When I began researching this piece, I would announce the phenomenon to friends. Minds were blown! Then I’d explain the science. Circuits were blown! The tune-out factor was real. The documentation is just pretty dense and inaccessible. A mere paragraph inside a contemporary research paper on the subject helps explain why DNA synthesis has yet to find its place in the popular imagination.
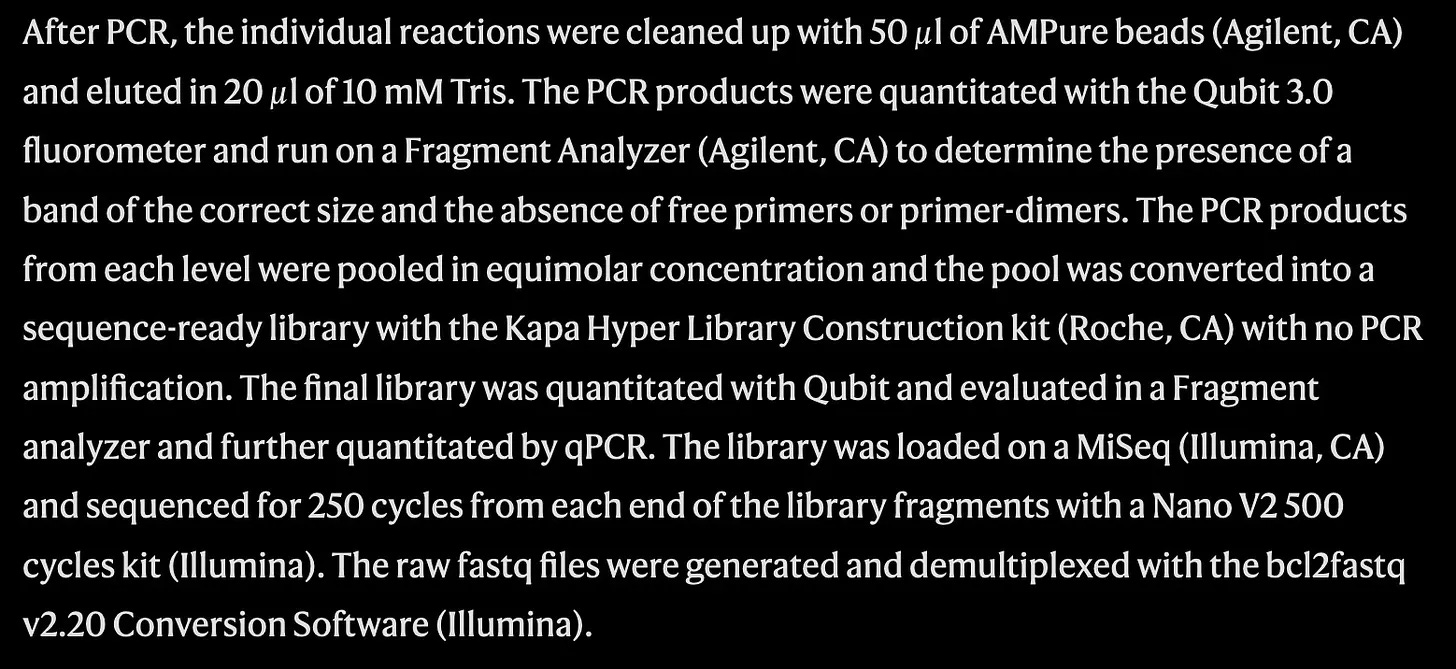

Riiiight. The question then is how to bridge the what and the how.
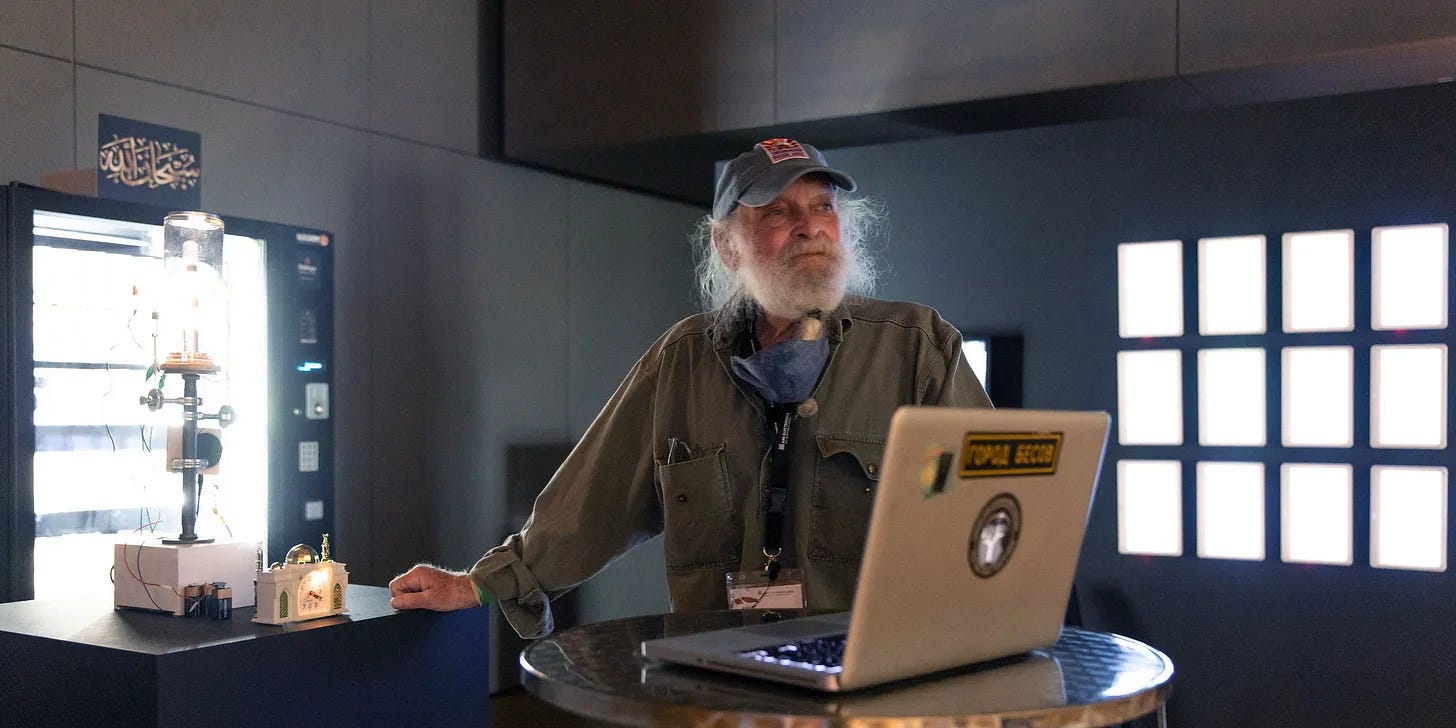
Going back to how we figured it all out in the first place is a serviceable path to technical understanding. My eureka moment came when I discovered a pioneering experiment conducted by a renegade bioartist named Joe Davis.
[Joe Davis Has Entered the Chat]
In 1988, Davis set out to write “extrabiological information” into DNA. His canary in the genomic datamine was a graphic icon of the Microvenus, an ancient Germanic rune representing life and “the female earth.” It looks like an ‘I’ superimposed on a ‘Y’. It also looks like something else…
To convert the graphic into DNA, Davis took inspiration from Carl Sagan’s ‘1079-Bit Message for Extraterrestrial Intelligence’, a graphic image converted to binary that was then inserted into a radio transmission and sent from the Arecibo Observatory in Puerto Rico to outer space.

The 2D Microvenus icon was translated into a 5x7 bitmap - an exact representation of the graphic using only ones and zeroes. It looked like this:
Here’s that binary strung out linearly:
10101011100010000100001000010000100
Davis’s next challenge was converting the 1s and 0s into the alphabetical characters representing the nucleotide bases in DNA: cytosine (C), thymine (T), adenine (A) and guanine (G). Nucleotide bases possess distinct molecular weights. Cytosine is the lightest, thymine is slightly heavier, which in turn is outweighed by adenine, and then the heaviest is guanine. Davis represented these weights with round numerical values accordingly:
C = 1, T = 2, A = 3, G = 4
The binary then just needed to be compressed into these numerical weightings. To accomplish this, Davis used a coding method now common in computer-compression technology. He assigned ‘phase-change’ values rather than incremental numerical values to the conversion. In this process, each DNA base (C, T, A or G) is represented by a binary sequence of 1s or 0s that are repeated until the base’s respective weight is hit. Once that triggers, the assignation process moves on to the next base. ‘10’ then creates the code for base ‘C’. If that sequence repeats, like ‘10101010’ you’d get ‘CCCC’. Throwing a ‘11’ or a ‘00’ at the end of that sequence would then code for ‘CCCCT’.
There’s isn’t a DNA conversion tool available online so I made a web app that converts text into binary then into DNA bases. You can check it out here.
ANYHOW, all told, the 35-digit Microvenus binary map above compressed to only 18 nucleobases:
CCCCCCAACGCGCGCGCGCT
Davis then chemically synthesized this DNA sequence into a form that could be inserted into a cell. This biological shipping container is also referred to as a vector. The vector Joe Davis used for his Microvenus experiment was a small extrachromosomal DNA molecule that can be absorbed by a cell membrane called a plasmid. Plasmids possess a built-in ability to recognize enzymes, which in turn serve as a filing cabinet for the encoded data. The Microvenus plasmid was cloned into a laboratory strain of E. coli at both Harvard and MIT in 1988. A sequencing or ‘reading back’ was performed successfully two years later.
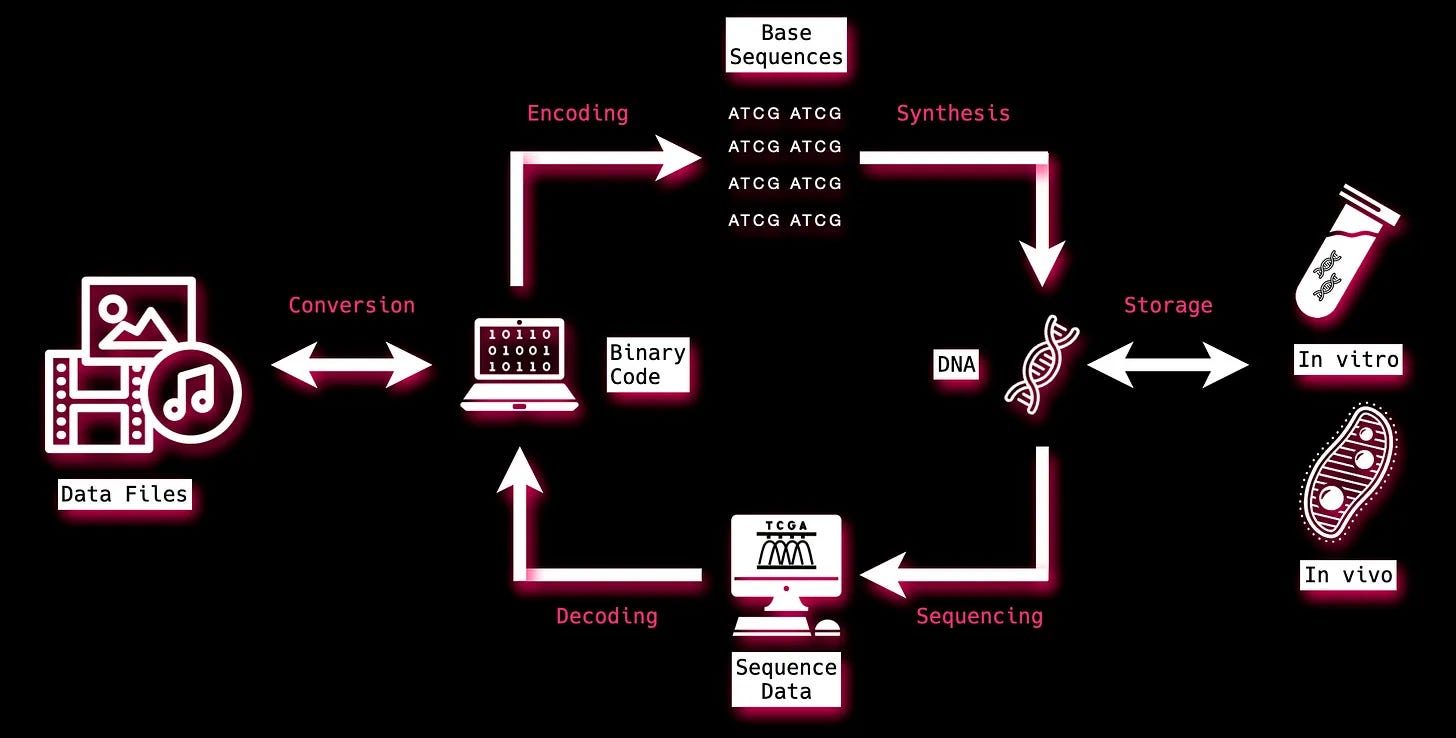
It’s been 44 years since Joe Davis’ Microvenus experiment, but the steps to insert data into and get data out of DNA remain roughly the same. Data is encoded from 1s and 0s to DNA bases. That information is then synthesized or ‘written’. Then it’s stored. When the data is needed, it is sequenced or ‘read’. Lastly, its decoded out of DNA bases and into binary bits again.
While the basic order of operations hasn’t changed much, the pace and scale of execution has been turbocharged by new organizations and shiny new tools. To get up-to-speed, here’s…
A Spotlight on Twist Bioscience
DNA’s emergence as a viable data storage solution has fueled an explosion in enterprise. Tech giants like Microsoft have wasted no time getting in the mix. The private sector is also teeming with smaller DNA storage-native startups. Austrian firm Kilobaser is developing the “Nespresso machine of DNA synthesis,” and Iridia is making DNA data storage available to communities and families. By far the most-discussed and well-funded DNA synthesis company is Twist Bioscience.
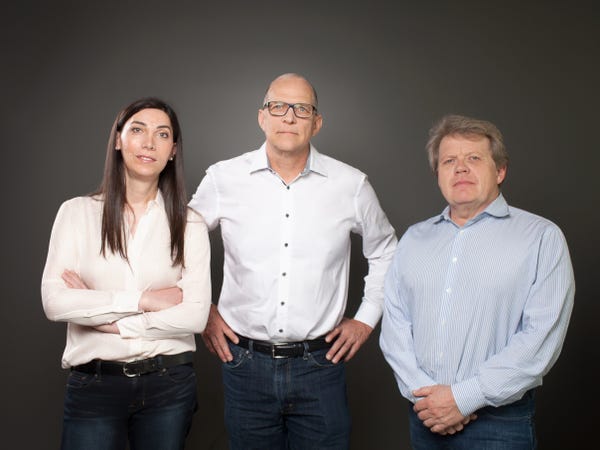
Twist was formed in 2013 by an applied chemistry dream team: Emily Leproust, Bill Banyai and Bill Peck (known as ‘The Bills’). Their combined backgrounds are uniquely suited to performing highly-intricate chemical acrobatics on a miniature silicon chip. CEO Leproust’s PhD in organic chemistry focused on microarray technology that enables thousands of small DNA snippets to be synthesized into genes simultaneously. Banyai brought a specialization in silicon micro-machinery to the table. Peck excelled at microfluidics, a multidisciplinary field in which microscopic amounts of liquid are transported around a silicon chip. Nine years later, Twist is a DNA synthesis powerhouse with 37 patents, over 1000 employees, and estimates over $200 million in annual revenue for fiscal 2022.
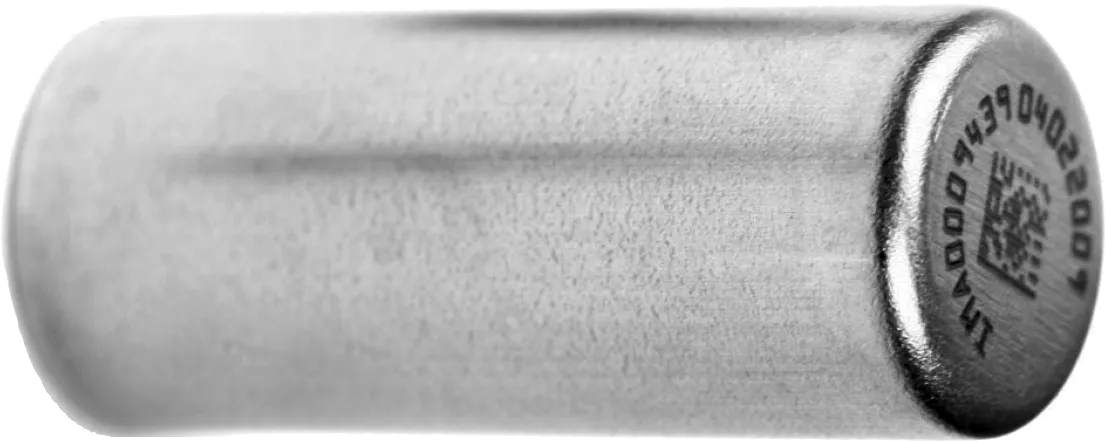
One of Twist’s many offerings is a frictionless e-commerce system that facilitates the purchase of a custom gene. A customer can go to Twist’s website, and enter a genetic sequence. Twist then verifies the sequence to ensure zero errors. While the gene manufacturing process begins, Twist conducts a biosecurity screen for both the customer and the requested gene. The whole process takes a few hours and costs $1,000 per megabyte of data. Your bespoke gene is then laser-sealed in a tiny vault called a DNA shell and sent off in the mail.
A Dimension Below
Twist’s premise from the outset was simple: make DNA synthesis smaller. By shrinking the hardware used to synthesize DNA, Twist aimed to parallel path more genetic combinations in a shorter period of time. Accomplishing this meant re-thinking the traditional lab set-up. Since the 1980s, DNA has been synthesized in 96-well plates - a collection of reservoirs about the size of two iPhones. The 96-well plate sort of resembles a high-tech muffin tin.
Each well accommodates 50 microliters, or about an eye-dropper’s worth of liquid. That solution is called a reagent and contains one oligo (short for oligonucleotide). Oligos are the key ingredient here. They are a short strand of synthetic DNA made up of a sequence of As, Ts, Cs, and Gs. Oligos serve as the basis for the expression of a custom gene. The synthesis of those 96 single-oligo wells guaranteed the production of one gene. It was all a very manual and time-consuming process.
Twist saw an opportunity to optimize the synthesis operation and produce more genes simultaneously. To do so, they upgraded the plates and wells of yesteryear with a silicon chip surfaced with a lot more micro ‘devices’ that could each accommodate an oligo.
“If you shrink that chemistry, you use 99.8% less reagents, and generate 99.8% less waste,” explained Angela Bitting, Twist’s SVP of Corporate Affairs. “In the same space where others make one gene, we make 10,000,” she pointed out.
Here’s a visualization of that chip, courtesy of Twist:
Twist’s “factory of the future,” as Bitting puts it, is about to cross another micro-spatial milestone. Heretofore, the chips Twist uses to make DNA space each oligo-holding device 50 microns apart. A forthcoming chip to be used for synthesizing DNA for data storage spaces these devices one micron apart.
Post-synthesis, the oligos go to a back-end process. Bitting broke it down: “It comes off the chip and it might go to the gene lab to become a gene, it might get aggregated into an oligo pool which are useful for things like CRISPR gene editing experiments, it might go to a critical workflow where you’re making a diagnostic to identify a rare disease, or those oligos could store digital data.”
Steffen Hellmold is Twist’s SVP of Business Development and a data storage industry veteran. Hellmold helped articulate how Twist’s huge strides in hardware miniaturization promise leaps forward in data storage. “If you want to store a megabyte of data, we can deliver that today,” Hellmold explained. This means it’s possible to write a megabyte of data into DNA with one pass of Twist’s machines without breaking the bank. “It’s not optimized for data storage, but it works,” Hellmold explained. The new Micron chip will take DNA data storage to the gigabyte class, meaning Twist’s writers will be able to insert a gigabyte of digital information into DNA in one pass for the same low cost. The next frontier is the terabyte class. “When this chip comes into existence in the not so distant future, we will surpass the highest density that has been delivered in the semiconductor memory to date,” pronounced Hellmold. “It is foreseeable that DNA data storage may become the driver for process technology in the semiconductor industry,” he predicted.

While an entirely DNA-powered data storage industry may be in our future, the technology isn’t ready for prime-time. Explaining where DNA data storage sits in its developmental arc, Director of Product Development Daniel Chadash invoked the Gartner Hype Cycle. “We’re in a place where Plasma TVs were in the late-90s,” Chadash explained. “It’s super-expensive and cost thousands of dollars but the technology works. You can write data, you can read data, you can decode it back to the file.” Chadash continued, “It’s possible to do end to end. Now it’s about scaling it and making it accessible.”
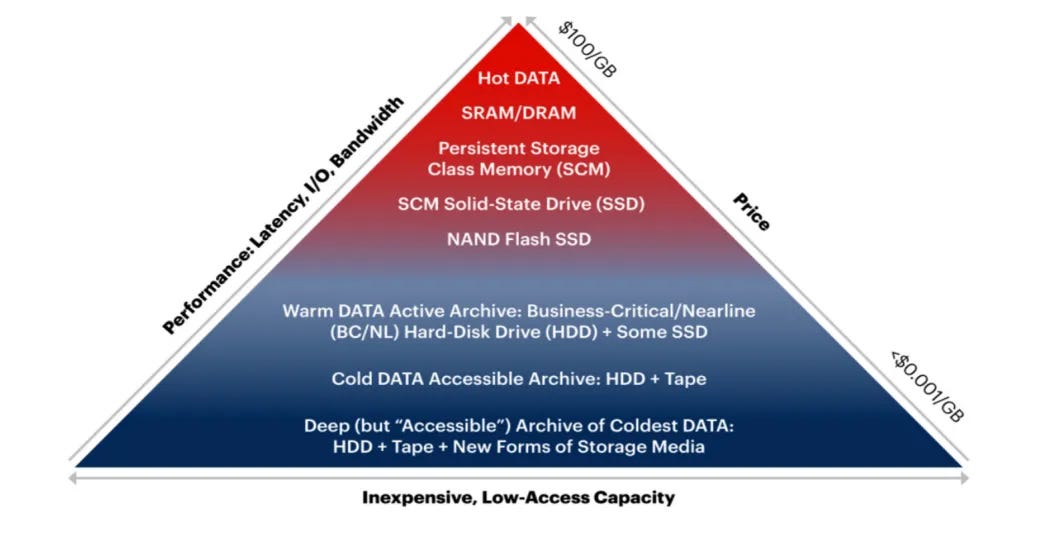
For now, DNA data storage is a tiny dot on the larger data storage landscape. A lens through which to make sense of this storage options jumble is the almighty dollar. The industry has a metric for this: Total Cost of Ownership (TCO). TCO is the amount of money it costs to store and maintain data. For this, the Storage Pyramid is useful:
While there are range of factors that contribute to a TCO, the lynchpin determinant of cost is the required frequency of access to the stored information. More access requires a storage solution that consumes more energy, requires more maintenance and is therefore more expensive.
It’s a Cold World
Storage accessed frequently is labeled ‘hot’. An example is autonomous vehicle sensor data, which companies need at the drop of a dime, all the time. That data is stored on high-performance drives that consume outsized energy. Storage solutions for data we access only semi-regularly, like yearly reports, is ‘warm’. Data we only need every once in a while, like archives, is cold and relatively cheap.

All told, cold storage comprises 60% - 80% of our civilization’s total info-stockpile. That percentage is growing, fast. Here’s why: As information gets older, our ongoing need for it decreases…but we expect continued access nonetheless. Consider the photos on your phone. The older a photo the less you revisit it, but that doesn’t mean you’re willing to delete it. That aging data, usually 90 days or older, all goes to cold storage.
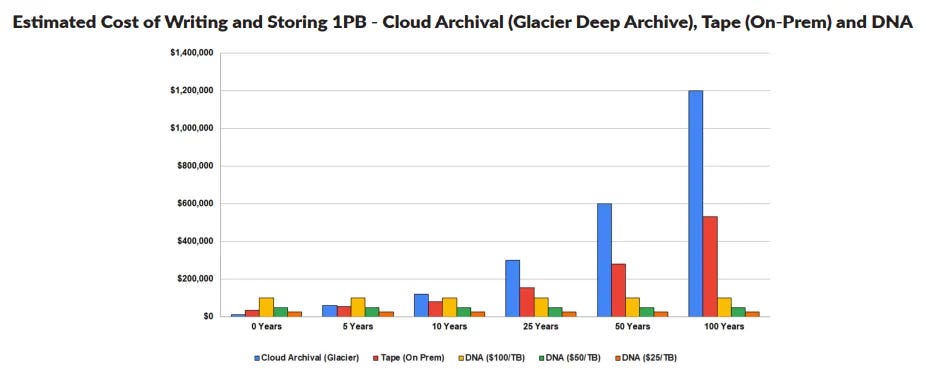
This ballooning of cold data storage is where DNA’s affordability, minimal space requirements and low maintenance needs pose the greatest upside. While near-term opportunities are around the corner, DNA’s longer-term cost benefits really start to pay off. The chart below projects the cost of storing one petabyte of data for 100 years using legacy storage solutions verses DNA. DNA’s cost-effectiveness in the long-term is unmistakable.
Strands of Fortune
Intelligence Advanced Research Projects Activity (IARPA), an organization within the Office of the Director of National Intelligence, predicts that the cost of synthesizing DNA by 2024 will be $1 per gigabyte; by 2030 it will cost $1 per terabyte. All in, total DNA data storage cost will reach ~$130 / gigabyte within the next few years. This affordability positions DNA very favorably within exploding cold storage space and the data storage industry at large.
Whether its DNA or another medium, the opportunity to fill a multi-zetabyte gap in the data storage market is big business. The storage market in total weighs in at $100 billion dollars per year. “Even if it’s only a subset at the bottom as cold storage data, it’s a multibillion dollar market,” Hellmold explained in reference to DNA. “This could easily be a multi-ten billion dollar market depending on how far DNA data storage moves up the stack,” he predicted.
Despite the staggering monetary opportunity or perhaps because of it, a spirit of cooperation prevails in the DNA data storage industry. Twist understands that inter-organizational cooperation is crucial to the technology’s adoption. This elevated perspective motivated Twist to co-found the DNA Data Storage Alliance. The Alliance is a union fait la force of organizations all invested in the success of DNA data storage. “One of the goals is to educate the market that this is no longer sci fi, but a reality that is coming true,” Chadash said. Unsurprisingly, biotech companies are quick to take a systems approach to achieve a healthy ecosystem. “The fact that we have competitors working side-by-side in this effort is essential to building a large-scale market,” Hellmold explained.
Stay Tuned!
The question remains: Can DNA data storage actually deliver on its promise of reducing the energy footprint of the data storage industry? In part 3, we explore the frontlines of this impact.